In a two-room laboratory sequestered in a hunkered-down building in Werribee, Victoria, a small but mighty group of baby frogs, some of the last bastions of their embattled species, are patiently waiting to die…
In this article Amalyah Hart takes a closer look at the reserch and technology which could be key to reversing the march of this species towards extinction.
This extended article is best suited for Year 4-12 Biology students who are interested in gene technology and species conservation. This could be used to teach about specific adaptations, natural selection and CRISPR – the powerful gene-editing technique which can be used to activate and edit genes.
Word Count: 5181

I’ve come to visit them on a blinding hot February day, and I’ve been excited about the encounter for weeks. But when I open the door and see them, squatting blithely in row upon row of plastic tanks, I’m struck with a potent
wave of tragedy.
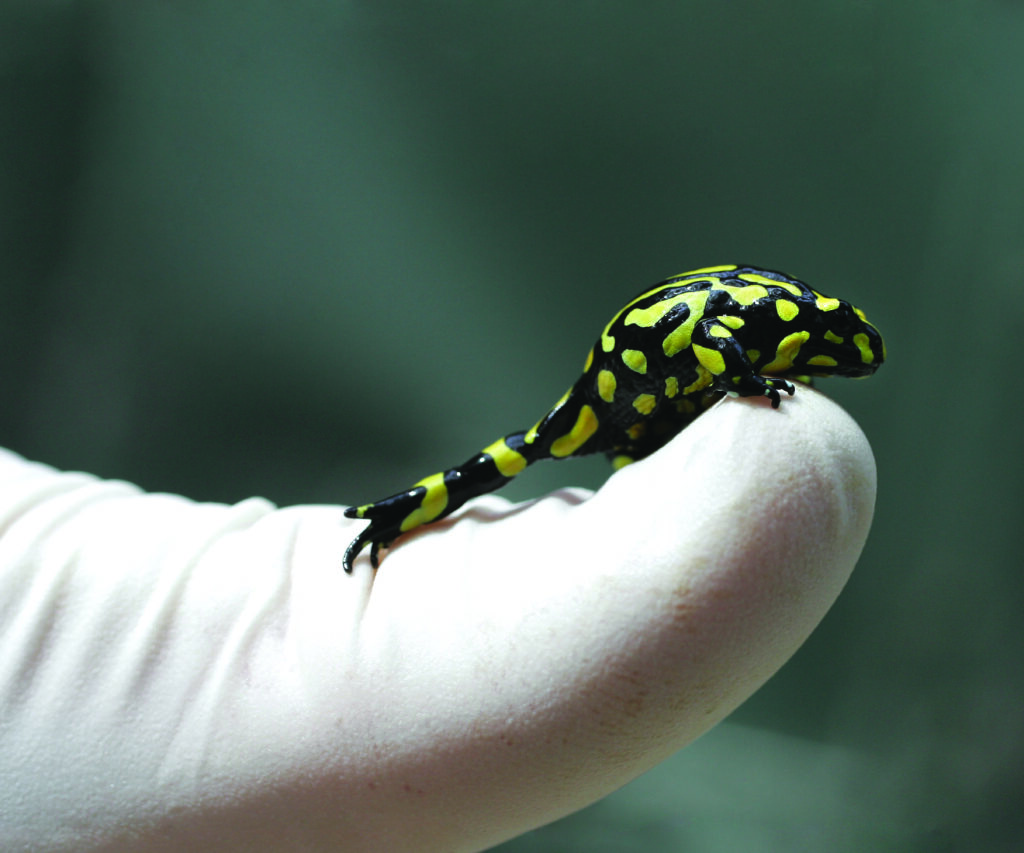
These are juvenile southern corroboree frogs, tiny little things with an overlaid pattern of bright yellow and black on their breathable skin, like a croaking nuclear waste sign. That’s pertinent, because they secrete a poisonous alkaloid through that skin that can kill prospective predators.
I squat on my haunches, greeting one of the little frogs as it hovers, suction-cup toe-pads pressed against the glass, its little throat moving rhythmically up and down. It has no idea what’s coming.
These young frogs, some no bigger than the tip of my thumb, are doomed – they won’t live for more than a few months at most. But their deaths may be the key to reversing the march of their species towards extinction.
That’s because these frogs are the first test subjects in a project that will plumb some of the most exciting (and controversial) realms of science, in the quest to conserve a dying species – by striding into the vanguard of gene editing.
Lee Berger was in the midst of her PhD at James Cook University in the 1990s when she began investigating an alarming and inexplicable global decline in amphibian populations that had been going on for at least 20 years. At the time, its effects were seen most acutely in the rainforests of Central America and Queensland.
Mass frog deaths were moving across the landscape, following the kinds of patterns you expect from an epidemic, and researchers were proposing that some sort of exotic pathogen must be behind the deaths. But it was Berger who, in 1998, first identified a fungus, Batrachochytrium dendrobatidis, suffusing the skin of sick frogs.
Southern corroboree frogs are native to the mossy sphagnum bogs of the northern Snowy Mountains, and are exclusively found within the limits of Kosciuszko National Park. They summer in mossy chambers, where the males voice a charmingly offbeat ‘squelch’ to attract their mates. In the winter, they retreat to the snow-sheltered undersides of snow-gum logs and leaf litter.
Southern corroborees are now functionally extinct in the wild, because B. dendrobatidis causes a ravaging disease known as chytridiomycosis. According to Zoos Victoria, there may be fewer than fifty of these frogs left in their natural habitat, and the tiny group remaining only persists because breeding programmes have periodically replenished their dwindling population.
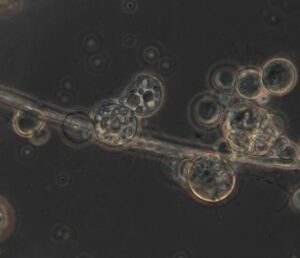
B. dendrobatidis, also known as the chytrid fungus, takes advantage of amphibians’ most important evolutionary quirk – their porous skin. “Frogs actually absorb oxygen through their skin, and they also absorb a lot of electrolytes that way,” explains Tiffany Kosch, a research fellow at the University of Melbourne and one of the lead architects behind a daring new plan to save the frogs.
“So, how the fungus ends up killing them is they actually have a heart attack, because they’re not able to maintain the correct electrolyte balance in their blood- stream, so the heart slows and they eventually die.”
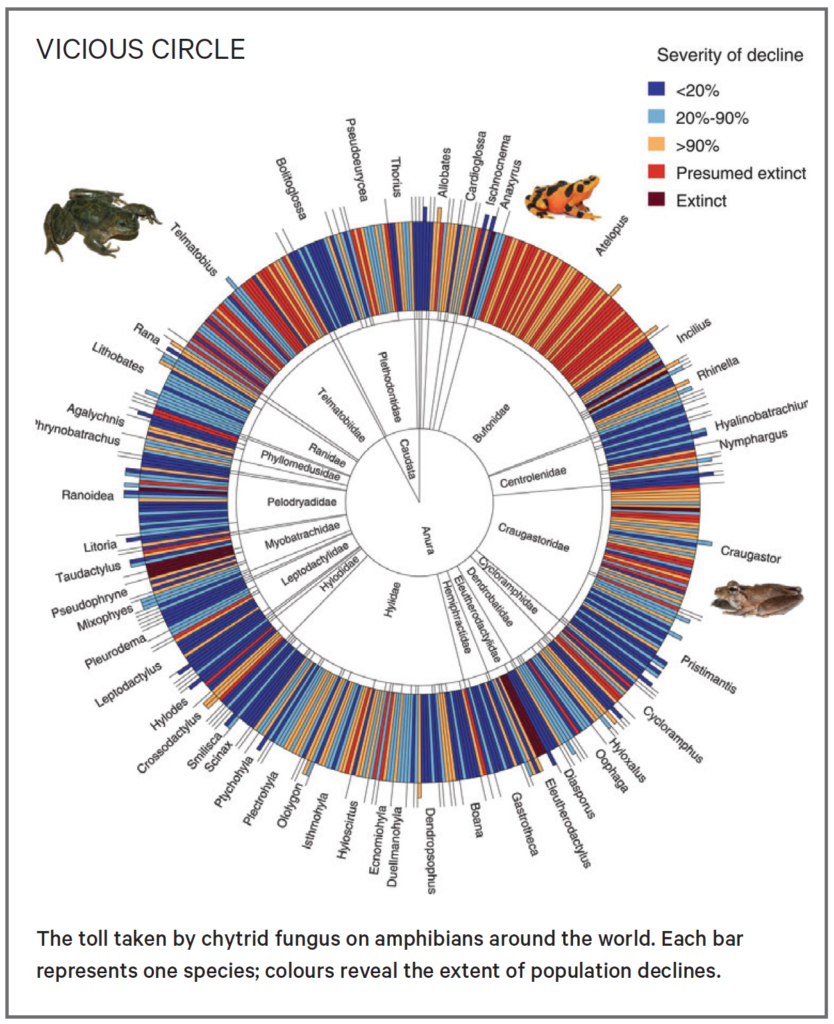
And the chytrid fungus is a serial killer: by some estimates, it may be responsible for the greatest disease-driven loss of biodiversity in recorded history. It travels in water, spreading through interlocking stream systems and from frog to frog, decimating the populations it comes into contact with.
In the Melbourne research lab, I’m introduced to chytrid by the scientist who first illuminated its crimes. More than 20 years on from her initial discovery, Berger is now working in the same facility as Kosch where, in a small fridge in the next-door room, she carefully stewards petri dishes of the murderous fungus.
She takes one out, places it under a microscope and invites me to look.

Tiny little swimmers dart around on invisible flagellae, dodging bigger, globule-like cells that hover, suspended in the artificial light.
The swimmers are the infective cells, which latch onto the frog’s skin and begin to multiply, transmuting into the fat globules that clog the frog’s life-giving pores. Looking at these tiny cells, floating inoffensively on the microscope’s plate, it’s hard to believe they could wreak such ecological havoc.
The chytrid fungus has resisted more than two decades of efforts to remove it, and the disease kills around 95% of the southern corroborees it comes into contact with. Kosch and her team at Melbourne University are aiming to reverse this seemingly intractable situation, by armouring the frogs with a set of genes that protects it.
This will require one of two approaches. The first option is artificial selection: breeding creatures with the necessary genes together to produce a resistant population over several generations. Humans have been practicing artificial selection for millennia, ever since the first domesticated crop or tamed wolf.
The second approach is a bit more controversial. Synthetic biology is an umbrella term for a number of techniques that manipulate genes or sets of genes to achieve a desired result.
These include transgenesis, which involves trans- ferring whole genes (or sets of genes) into one species from another species, and gene-editing, which involves ‘snipping’ out certain genes (or parts of a gene) and replacing them with others.
Synthetic biology can alter an organism’s genotype (its genetic material) to produce a desired phenotype (the observable traits coded for by the genotype).
“So many of these frogs are now bred in captivity, but whenever they release the frogs they usually don’t survive, because the pathogen can’t be eradicated,” says Kosch. “So, our idea is to test synthetic biology methods, which are very successful in agriculture, and see if they might work for conservation.”
More: The Future of Food
About 10,000 years ago, as the last ice age relinquished its grip and H. sapiens stood on the threshold of the Neolithic, archaeologists think that a handful of communities living between the Tigris and Euphrates rivers in Mesopotamia (modern day Iran, Iraq and Syria) began to play around with the seeds of wild plants, purposefully selecting and planting them for a predictable crop, and the certainty of food. In doing so, they could not have known that their selective choices would alter the genomes of entire species.
The agricultural revolution completely reshaped the structure of human life, enabling more perma- nent settlements and larger communities. And the crop plants that would emerge became both genet- ically and phenotypically distinct from their wild counterparts.
Successive civilisations have been meddling with DNA for millennia. What makes synthetic biology so different?
Current gene editing techniques allow scientists to make the kind of changes in a single generation that might once have taken many iterations of environ- mental trial-and-error. Proponents argue they also allow for much more precise changes.
“The really good thing about gene editing, compared to artificial selection, is you’re only changing the genetic elements that you’re trying to change, not a whole bunch of other things,” explains Kosch.
“And that’s really important, because we want these frogs to be able to survive in the wild in future.
And because we don’t know what these frogs are going to be experiencing in the future, we have to preserve their genetic diversity.”
But that also makes it an impressively powerful tool, in the hands of a species that has not always been judicious.
Whenever you have a technology that’s really, really powerful, you can use it to do really good things, but you might use it to make big mistakes, or some people might use it with bad intentions,” says Christopher Gyngell, a bio-medical ethicist at the Murdoch Children’s Research Institute who has worked extensively on the ethics of gene editing.
That’s the crux of this issue, we’ve got this power now, and we have to decide how to use it.
For Gyngell, the important thing in this type of research is to be slow, cautious, and reasoned.
“I think you need to be careful with these technol- ogies, you need to move slowly,” he says. “What human history has shown us is that ecosystems are really complex, and sometimes humans don’t understand them as well as we think we did.”
One of the core criticisms of using gene-editing in conservation, according to Kosch, is the idea that it’s a band-aid over the problem, and that scientists and conservationists should instead focus on eradicating the threat.
The problem is that many of the threats to biodiversity around the world are now rampant – a Pandora’s box of dire consequences.
Limiting global warming now won’t stop or reverse its consequences for decades, at best. It’s the same with chytridiomycosis.
Scientists have tried for decades to come up with ways to eradicate the disease, including expensive and dangerous chemical treatments with knock-on effects for other local species, and all to no avail.
From Kosch’s perspective, then, the way to give this frog a fighting chance is to delve into its genes.
Only a few weeks after my first visit to the lab, I receive an email from Kosch: the experiment is underway.
The researchers have placed the frogs in little takeaway sauce cups. After pipetting a liquid mixture of water and chytrid fungus over them, they’re kept in the containers for six hours, to allow the fungus to fully take hold. Then, over the course of the next few weeks, the team will observe how each frog is faring.
Kosch, like any good biologist, loves frogs, and she admits it’s hard work to do. But she is focussed on the research she believes is the species’ best hope of survival.
“Obviously we don’t want to be causing frogs to die, but at the moment unfortunately a lot of these little guys are not going to survive if they’re released into the wild,” she says. “If we’re doing something that can maybe increase their chances of survival someday, it’s worth trying.”
Around 95% of the frogs will die within a few weeks of infection. The first indication that something is wrong is a kind of limpid sluggishness, a reluctance to move. The scientists put a gloved hand into the tank: a healthy frog will hop away, but a frog sick with chytridiomycosis will stay put.
Another tell-tale sign is posture: a healthy frog will sit with its legs tucked underneath it, whereas an unhealthy frog will splay its legs and struggle to climb.
In the final test, the researchers will take the frogs out of their tanks and place them upside down in the palm of their hand. A healthy frog will right itself, but a sick frog will lie there, helplessly supine. After this righting reflex is tested and failed three times, the frog will be euthanised.
The hope is that 5% of the frogs will survive infection and recover, and their DNA could hold the key.
By looking closely at the genomes of the survivor frogs, the team hope to identify the genes that are affording them resistance to the pathogen.
“This might be hundreds of genes, maybe thousands, or it may just be a few,” notes Kosch. “At this point, we really don’t know.”
But once they do know, the discovery will allow the researchers to work out how to arm new gener- ations with the right genes that could help them survive in the wild.
So, do they use the old method or the new? Artifi- cial selection or synthetic biology?
If the genes that code for resistance in these frogs are easily inherited, or involve many genes all of which have a small effect, they’re likely to pursue selective breeding, because it may be easier to breed the genes into a new population than to snip and replace so many, low-impact genes.
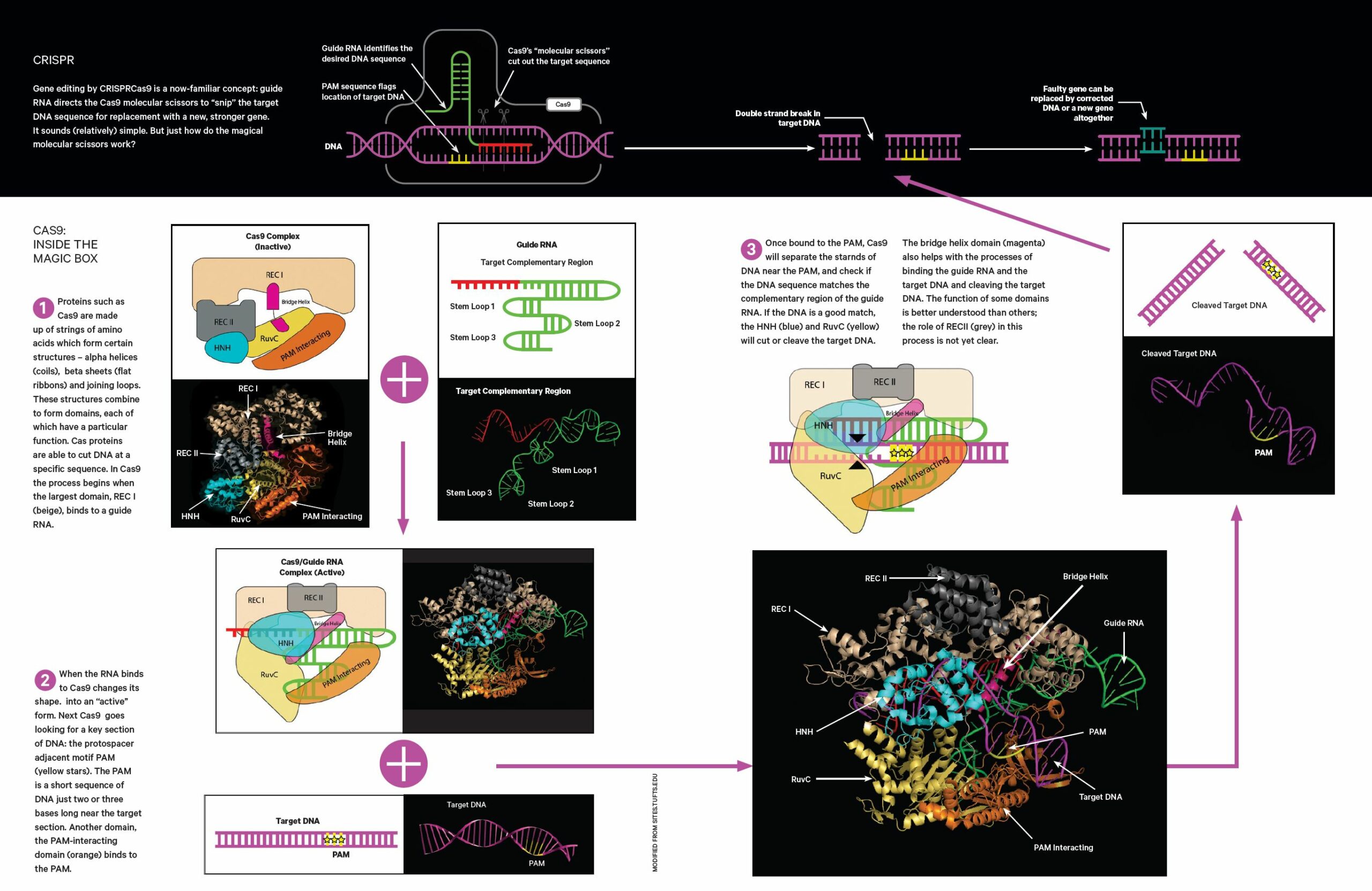
If the genes have low heritability, or they involve fewer genes that each have a significant effect, Kosch says they’ll likely opt for gene editing, using CRISPR-Cas9 (see following pages).
That’s because, in this case, it would be more diffi- cult to breed a resilient population, and also because gene editing is much quicker – artificial selection can take tens or even hundreds of generations for the positive genetic change to take hold.
Both methods have risks: selective breeding a small population of can reduce genetic diversity in the population, or could breed out certain genes needed to survive in the wild. Inserting genes could carry markers for chytrid resistance, but also insert another unforeseen weakness.
And there’s another problem: releasing genetically engineered frogs into the wild is uncharted territory, so the methods they use will have to adapt as the regulatory landscape around this strange new world catches up and starts to take shape.
The chestnut tree is an icon of Americana; you’ll have heard the Christmas song about roasting its nuts on an open fire. The American Chestnut tree was also a cornerstone species for local ecosystems up and down the US east coast for millennia.
“This tree was a very abundant mass producer,” explains William Powell, director of the American Chestnut Research and Restoration (ACRR) project at the SUNY College of Environmental Science and Forestry, New York, US.
“It produced a lot of nuts for wildlife [and humans] to eat, it produced really straight-grained, rot-resistant wood for people, and the leaves were used as medicine by Native Americans as well as early settlers.”
It’s a tree that provokes emotion for many of the people who live in its native homelands. “It’s ingrained into our folklore,” Powell says.
But the chestnut has been all but lost to the east coast, thanks to another invasive fungal marauder– Cryphonectria parasitica, or chestnut blight. First observed on US soil in 1904 in the New York Zoological Gardens, the blight was introduced when people began importing the Asian chestnut tree into the country.
“They didn’t know at that time that when you bring a tree over, you bring all its microbes over also,” Powell says. While the fungus was a mildly annoying skin condition for the Asian chestnut tree, it spelled disaster for its American counterpart.
The fungus latches onto the tree’s bark and starts to gnaw a wound, called a canker, into the trunk or branch. It does this by producing a substance called oxalic acid, which kills off the tissue in front of it to produce a necrotising substance that the fungus can eat.
The canker grows, eventually girdling the trunk or limb and cutting off the flow of nutrients; everything above the canker dies. If a canker forms at the base of the tree, everything above ground perishes.
It’s estimated that over the last century, four billion American chestnut trees have disappeared because of the fungus, with small pockets remaining in the Carolinas, West Virginia and in Pennsylvania.
Chestnut trees can survive at the roots, so there’s still a few million stump sprouts left. Periodically these stumps grow, encounter the fungus and die back, so the viability of a flourishing chestnut tree along the spine of the east coast is low.
But the existence of these stump sprouts means there’s a relative wealth of chestnut genetic diversity, so conservationists seeking to breed a new (and perhaps improved) population aren’t hampered by the problem that an endangered species tends to have: low variability in the population genome.
That’s where Powell’s research group comes in. For several decades now, Powell and his colleagues at the ACRR have been using genetic engineering techniques to alter American chestnut tree cells in the lab, with the goal of creating a tree that’s resilient to the fungus – and it seems to be working.
They’re utilising transgenesis, with a nifty little gene borrowed from the wheat plant. The gene produces an enzyme, oxalate oxidase, that detoxifies oxalic acid, breaking it down into carbon dioxide and hydrogen peroxide, two compounds the plant can use.
“And so, the nice thing is it doesn’t actually hurt the fungus at all,” says Powell – who is perhaps more charitable than me. “All we’re doing is taking the weapon away, so now it can live as a saprophyte [decomposer] on the tree, it can still cause a little bit of damage, but it’s not the severe damage where you get the whole tree being killed.”
They’ve transferred the gene into the tree cells using another handy bacterial tool, agrobacterium-mediated transformation.
“Agrobacterium is a bacteria that naturally genetically engineers trees or plants in general, and has been doing that for millennia,” says Powell. “So that’s what we did with the oxalate oxidase, we made what’s called a vector, put it into the agrobacterium, and then let the agrobacterium put it into the tree.”
Once the gene has been introduced, the researchers kill off the bacteria with antibiotics. This is all done while the tree is still in its cellular stage in the lab, so it’s tightly controlled.
The team have been field-planting these trees since 2014. But the next step – actually releasing them into the wild and populating the whole coast with its beloved chestnut once more – could take years of further regulatory hurdling.
“Those [field-planted trees] are all planted under USDA permits,” explains Powell. “And they come out and inspect our fields, we have certain rules to follow, and we can’t let them pollinate right now.”
At the moment, the program is under review by three agencies: the US Department of Agriculture (USDA), the Environmental Protection Agency (EPA), and the Food and Drug Administration (FDA).
Reviews for transgenic plants in the past in the US have almost exclusively focused on agricultural plants that are harvested every year; the chestnut, on the other hand, is not only a wild-dwelling plant but a long-lived tree.
The concept of changing an organism’s genes and then releasing it back into the wild is intrinsically Frankenstein-ish to some. Ever since scientists carefully unpicked the double-helix 70 years ago, DNA has been something of a godhead – an inviolable biological structure.
But that’s not always how scientists see it.
“Part of the root of some scepticism is that there’s something kind of integral and wholesome about a genome, and that that kind of defines what a being is,” says Andy Newhouse, an ecologist and assistant director of the project at SUNY.
“But as we’ve been learning more and more over the past couple of decades about where our genes come from, they get swapped around all the time.”
Some of this resistance may stem from a perspec- tive that humans and other creatures are all separate entities, rather than one member of an interactive community or organisms.
“There are so many genes in people, in plants, in chestnut trees and in corn from other stuff; from viruses, from bacteria, from related plants and unrelated plants,” Newhouse points out. “So, I wish I could help convey that the genome doesn’t define what a being is.”
So, what do Powell and Newhouse make of fears about Franken-tree?
“If you actually look at this kind of agrobacte- rium transformation compared to older, traditional breeding methods like hybrid breeding, it turns out this agrobacterium transformation causes ten-fold or maybe even 100-fold fewer changes to the genome than all those older techniques,” says Powell.
“And so, in reality, especially for conservation where you’re trying to preserve the identity of certain species, it’s kind of nice to have less of those changes.”
The next time I visit the Werribee frog lab, there’s a chill in the air. I slip on a pair of gum boots and a lab coat at the door so I don’t unwittingly track the fungus out of the room and commit biological warfare against other frogs in the building.
Of the 58 frogs I met before, just four are left – their lonely plastic tanks dwarfed by the empty shelf space where their comrades once sat. But there’s a glimmer of hope to this otherwise sad story, because something quite remarkable seems to have happened.
One of the frogs – tank 27 – has escaped the inevitable. This frog doesn’t seem to have contracted the disease at all, despite sitting in a liquid mixture of live chytrid fungus for six hours.
Given that Kosch only set out to identify frogs that could survive infection, the prospect of a frog that could actually be resistant to the infection itself is tantalising.
“We don’t know if it’s an anomaly or a super resistant frog, but it’s a weird little thing,” she says.
It’s possible that somehow this particular frog was given a quantity of dead fungus – but Kosch says that’s unlikely. The next phase of the study, then, will show them whether any other frogs have this mysterious super-resistance.
Apart from super-frog and his three dying friends, the other 54 all succumbed to the bug, and were euthanised when they failed the final reflex test. But Kosch and colleagues have painstakingly preserved tissue from every last one.
“We’ve kept the skin sections and we’re going to study the skin and see if there’s anything unique,” she says. “Then, we’re going to actually develop a genotyping method from the tissues of these frogs, so not a piece of them will be wasted.”
Genotyping is a technique that can detect mutations in DNA between individuals of the same species that can lead to major changes in the phenotype. Genotyping will be crucial in identifying which genetic mutations the resistant frogs share that might be transferrable.
There are all sorts of methods for genotyping, but the one the team hopes to develop using the pilot- study frog tissue with another frontline tool.
“The most common method for wildlife is really crude, you basically just digest the genome with what are known as restriction enzymes, and then you look at certain sized pieces that remain and you sequence those to look for variation,” Kosch explains. “So, you don’t really have any control over what parts of the genome you look at.”
SNP-ChIP is a bit different. SNP stands for single nucleotide polymorphism, which describes a change in the genetic code at the level of a single base pair on the genome – essentially the fundamental unit of the DNA double-helix. Base pairs contain a combination of two of the four nucleobases found in DNA: either adenine and thymine, or guanine and cytosine and the rungs on the double helix ladder.
If a base pair is polymorphic, some members of a species may have that base pair at that location on the genome, while others will have a different base pair. There are only two potential base pairs in DNA: adenine pairs with thymine, and. It helps if you imagine the genome as a long metal chain: ten links into the chain some people might have a gold link, while others may have a silver link.
So, SNP changes are very small but very common – on average, an SNP occurs after every 1,000 base pairs in the genome (for reference, the human genome contains approximately 3 billion base pairs, while the Southern Corroboree frog genome contains three times as many).
The SNP method is thus more refined than other traditional genotyping methods
To identify which SNP a strand of DNA has, scientists will introduce synthetic, fluorescent nucleobases that bind to the corresponding bases to flag potential genetic variations.
If the frogs that survive chytridiomycosis share some clear SNPs, that will signal to the research team where their hunt should focus. If those SNPs are close to genes that are known in other species to code for resistance to other pathogens, or sit close to genes known to code for an immune molecule like a T cell, that’s another potential target.
Since there will likely be at least 50,000 SNPs per frog, they’ll need a computer program to conduct statistical analysis to identify potential candidates. And even then, it’s not perfect.
“There will be some [SNPs] that are found in the resistant frogs that have nothing to do with it, they just happen to be there,” Kosch says. “So then you’ve got to try and rule that out as well.”
It’s going to be a complicated and painstaking process of elimination. But if they find that mutation needle in the DNA haystack, they might be able to save the species.
Whether you’re nervous about Franken-frog or not, this kind of work is likely to become more common, as we enter the adaptive phase of our reckoning with the compounding impacts of climate change, habitat loss, and biodiversity collapse.
A million species are at risk of extinction, according to the UN. Most of these are disappearing because the equilibrium they evolved under is shifting. Oceans are warming, tree canopies are being replaced by arid grasses, rainfall patterns are growing or shrinking, fires are breaking their historic bounds. Change is coming. In many cases, change is already here.
So, do we owe it to the species we’ve imperilled to save them, by whatever means necessary? Or is this just another experiment in human hubris?
“In the face of more and more pressing threats, from climate, invasive species, other biodiversity threats, using all the powerful tools, including genetic engineering, is necessary,” argues Newhouse.
Gyngell, within reason, tends to agree.
“My perspective on it is very technology neutral,” he says, “so I don’t think technologies are good or bad, it’s how you use them.
“I think we can just have this mentality that we need to protect our ecosystems, and we should be using whatever tools we’ve got at our disposal to do that.”
For the southern corroboree, which evolved in tune with its now threatened homelands on the Snowy Mountains – an area that was almost wiped out by the 2019-20 bushfires – things are looking dire. Without a pair of molecular scissors, argues Kosch, we might not be able to save them.
“If we want these frogs to be back in the ecosystem this is the only approach that has plausibility
This article was written by Amalyah Hart, for Cosmos Magazine Issue 95.
AMALYAH HART is a UK-based graduate of archaeology and anthropology and a former Cosmos journalist. Her previous story –on bio-inspired materials – appeared in Issue 90.
Cosmos magazine is Australia’s only dedicated print science publication. Subscribe here to get your quarterly fill of the best Science of Everything, from the chemistry of fireworks to cutting-edge Australian innovation.
Login or Sign up for FREE to download the educational resources